
Complete Genome Sequence of SMBL-WEM22, a Halotolerant Strain of Kosakonia cowanii Isolated from Hong Kong Seawater | Microbiology Resource Announcements
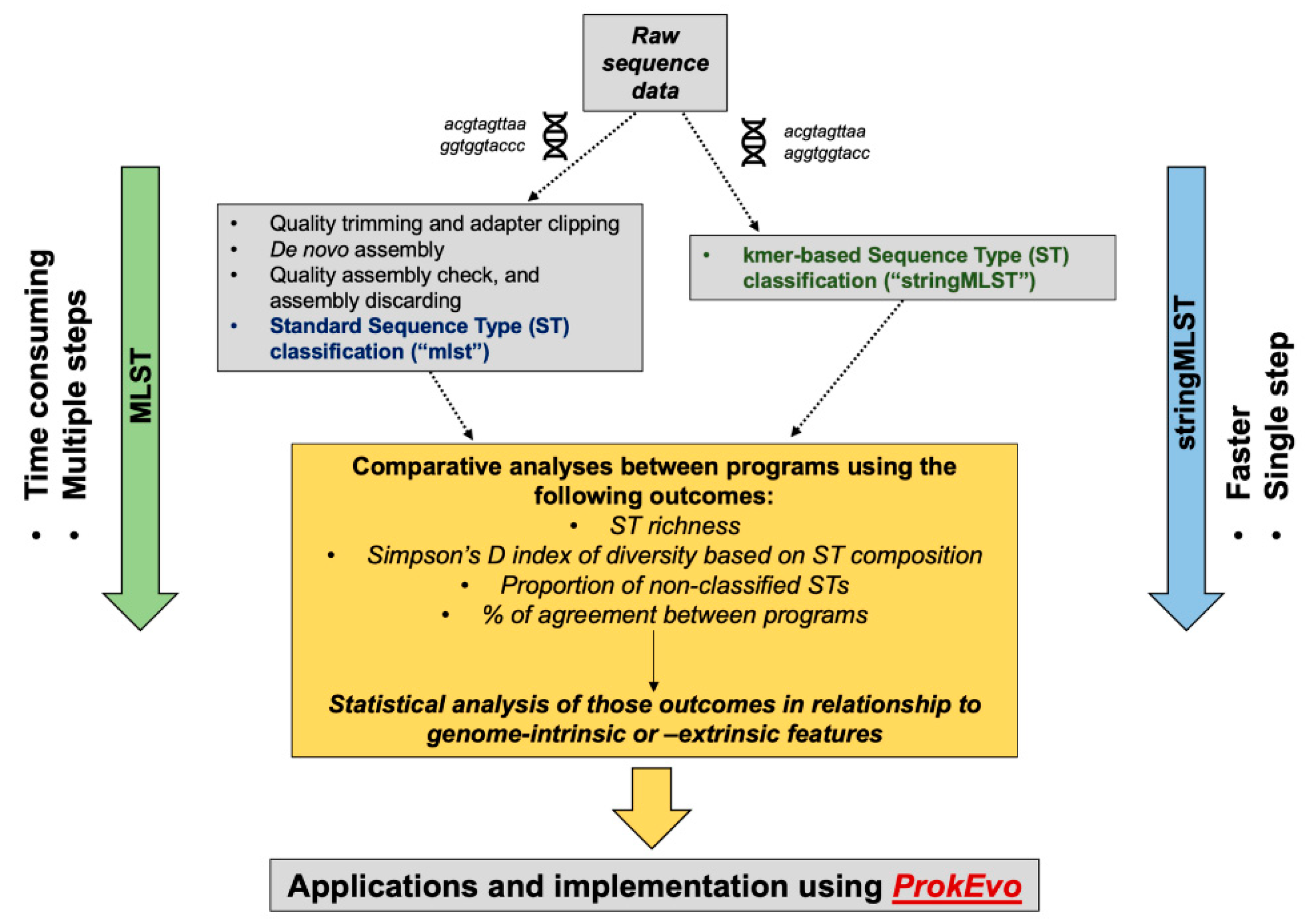
Life | Free Full-Text | Systems-Based Approach for Optimization of Assembly-Free Bacterial MLST Mapping | HTML

Chromosome-scale and haplotype-resolved genome assembly of a tetraploid potato cultivar | Nature Genetics

Draft Genome Sequence of Streptomyces sp. Strain FB2, Isolated from Rice Rhizosphere | Microbiology Resource Announcements
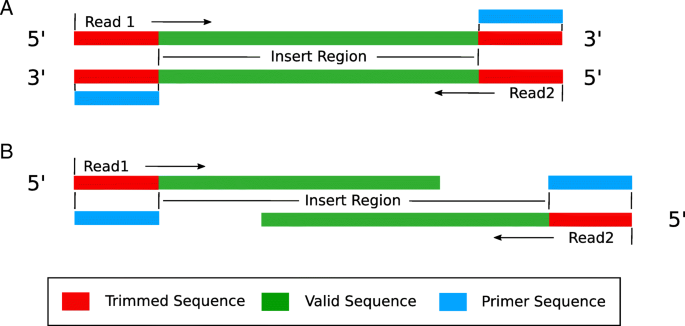
pTrimmer: An efficient tool to trim primers of multiplex deep sequencing data | BMC Bioinformatics | Full Text

Complementary combination of multiplex high‐throughput DNA sequencing for molecular phylogeny - Suyama - 2022 - Ecological Research - Wiley Online Library

HTSQualC is a Flexible and One-Step Quality Control Software for High-throughput Sequencing Data Analysis | bioRxiv

Microbiome Analysis Laboratory, CNB-CSIC on Twitter: "We are proud to know that FISABIO (Public Health Reasearch Services in Comunidad Valenciana, Spain) has adopted #SqueezeMeta as its reference tool for metagenomic analysis, and #

A modified CUT and RUN protocol and analysis pipeline to identify transcription factor binding sites in human cell lines: STAR Protocols
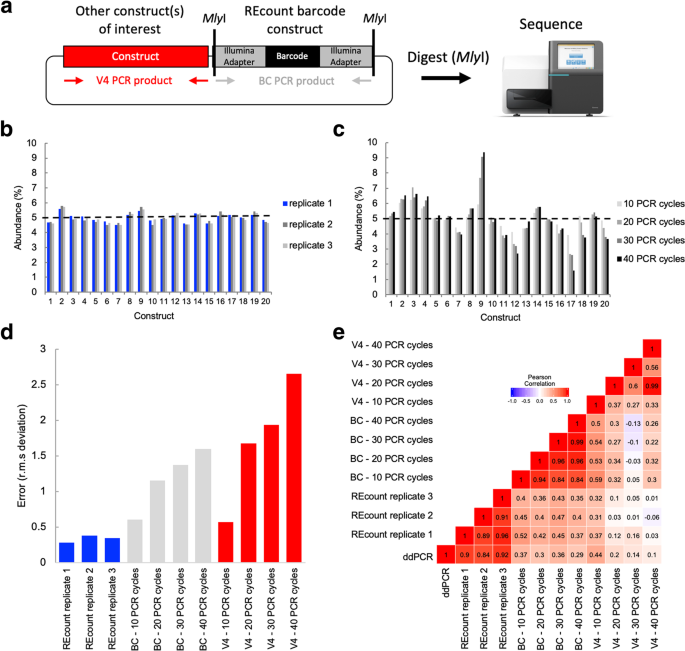
Measuring sequencer size bias using REcount: a novel method for highly accurate Illumina sequencing-based quantification | Genome Biology | Full Text


![PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc90910b6e31fe44cddc1e341f21eec0aaa5db44/6-Table2-1.png)

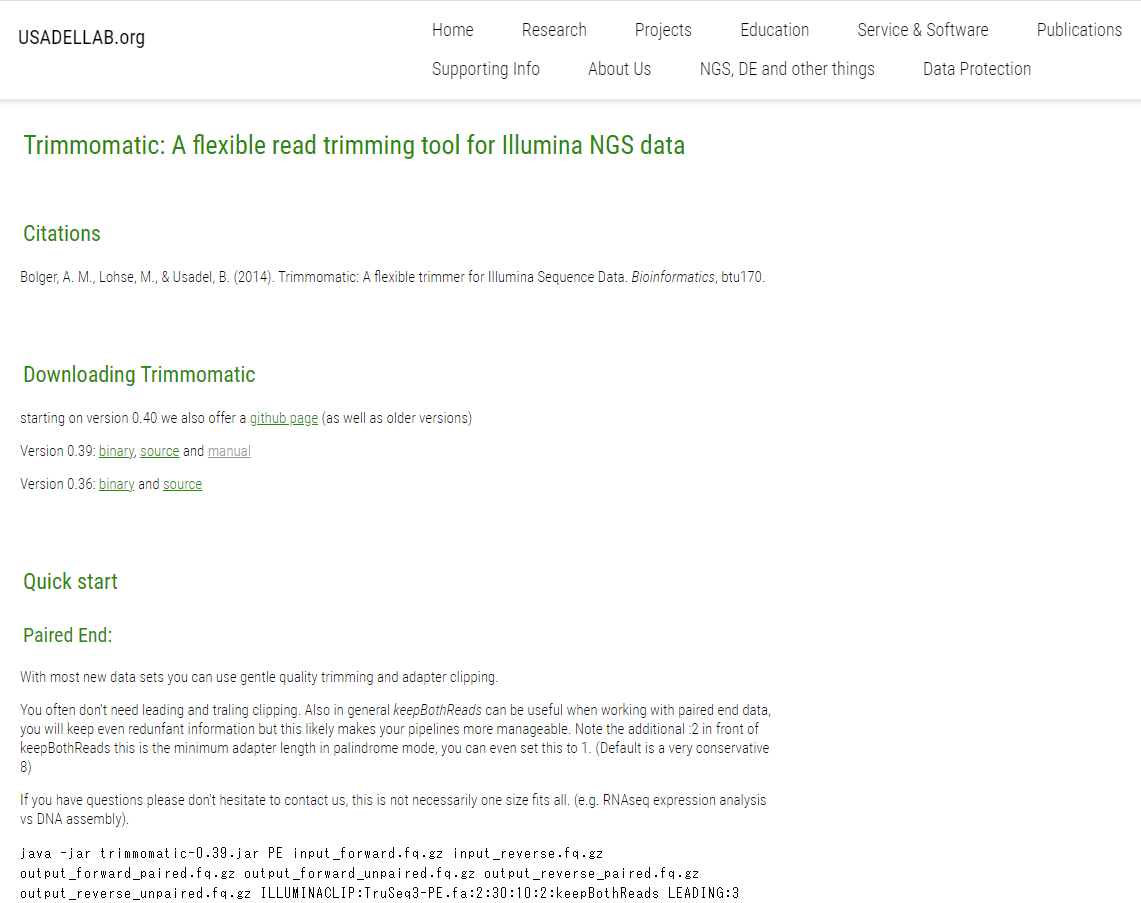





![PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc90910b6e31fe44cddc1e341f21eec0aaa5db44/2-Figure2-1.png)


